Summary
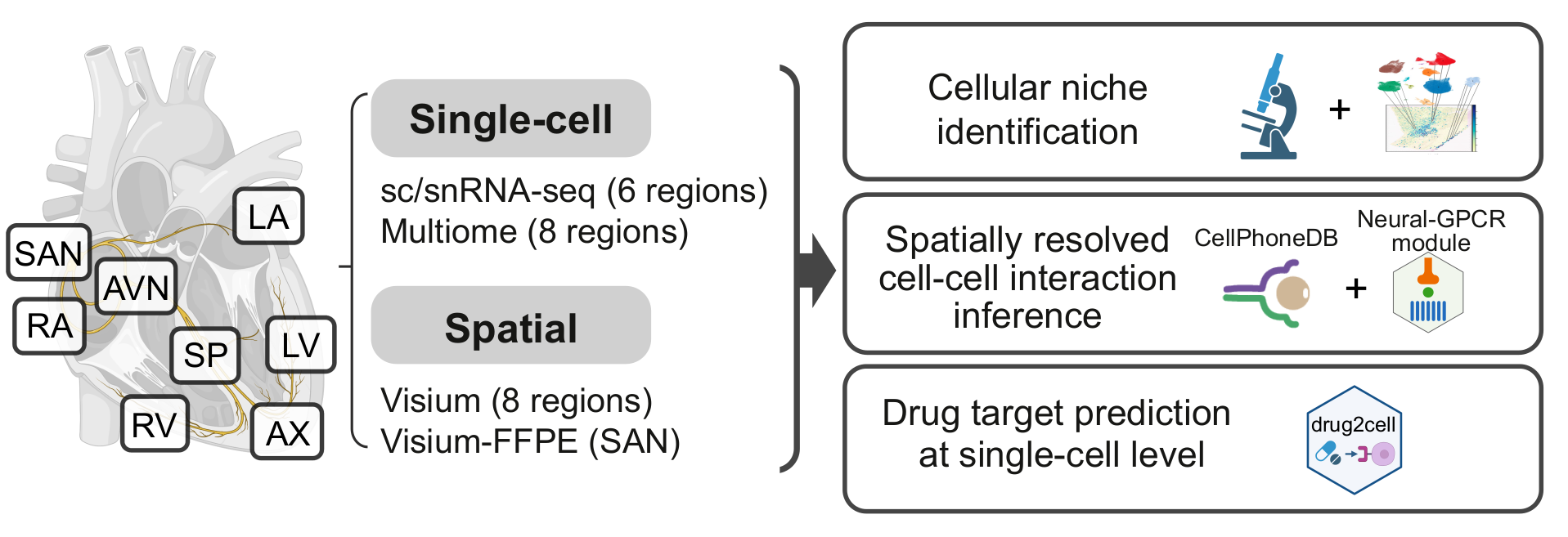
Using single cell technologies, we profiled the transcriptome of 704,296 cells and nuclei eight anatomical cardiac regions including the cardiac conduction system: left and right ventricular free walls (LV and RV), left and right atria (LA and RA), apex (AX), interventricular septum (SP), sino-atrial node (SAN) and atrioventricular node (AVN). Using state-of-the-art machine learning methods, we identified 12 major cardiac cell types and 75 different cell states.
Cell states defined by sc/snRNA-seq data were mapped in spatial transcriptomics data from the eight regions using cell2location.
Raw reads for the newly generated data are available in ArrayExpress here: Multiome-RNA, Multiome-ATAC, and Visium.
Donor demographics
The data consists of healthy hearts from 25 adult donors, 14 female and 11 male, within the age range of 20 to 75 years old.
All tissue was from transplant donors without a history of cardiac disease or arrhythmia and hearts contributing to the SAN and AVN regions were from donors with normal conduction parameters confirmed on 12-lead electrocardiograms prior to donation.
Cardiac conduction system
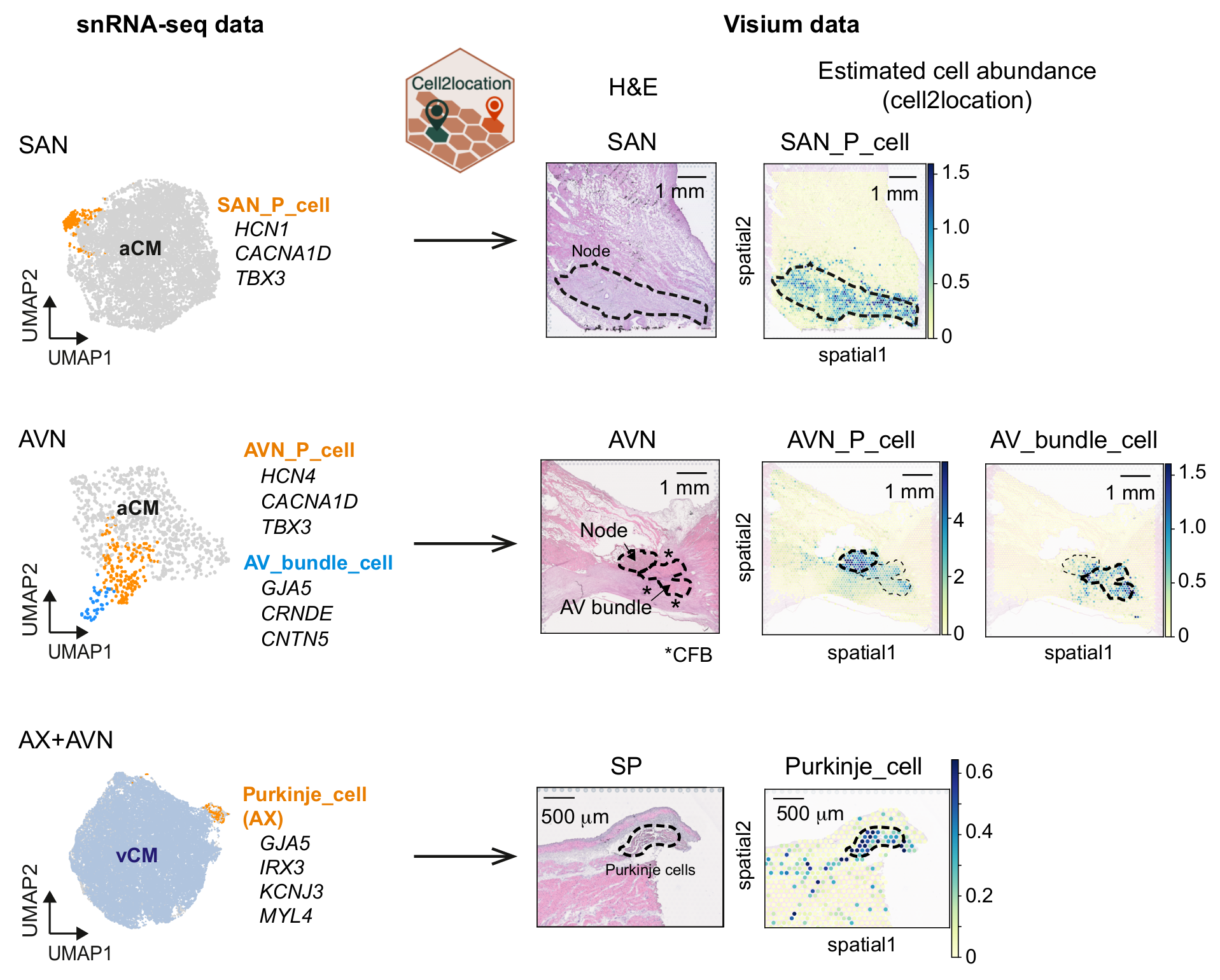
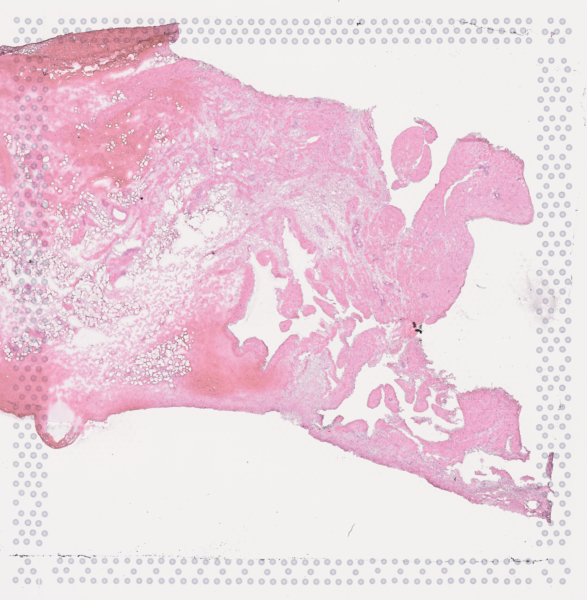
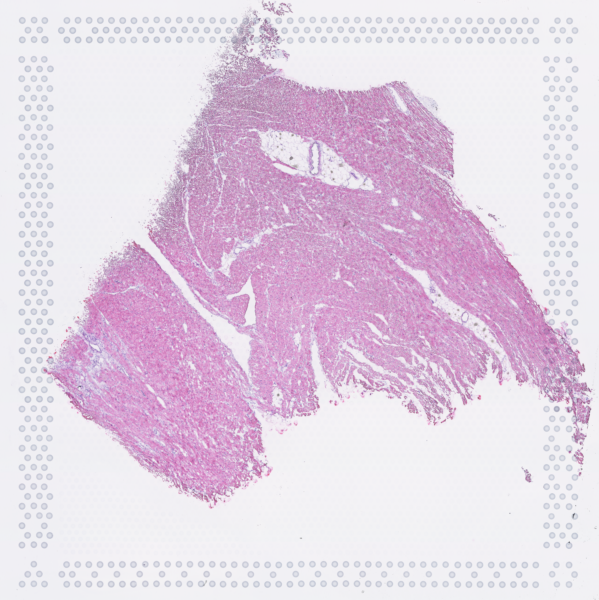
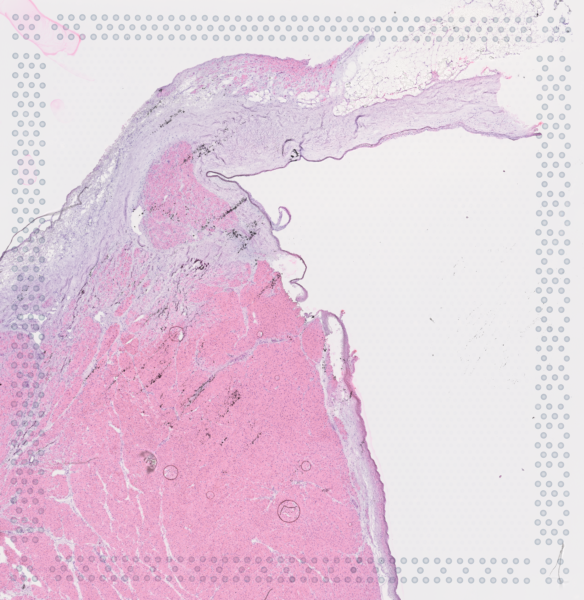
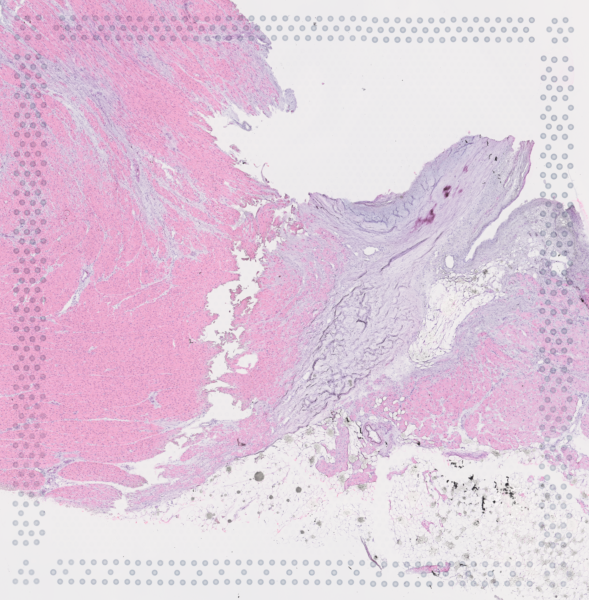
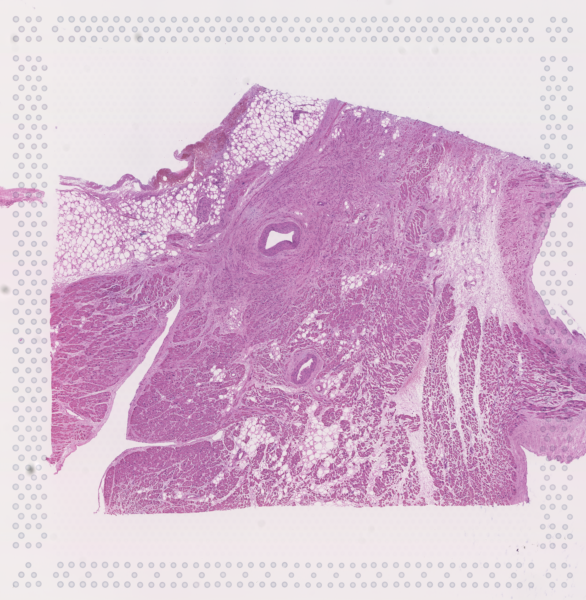
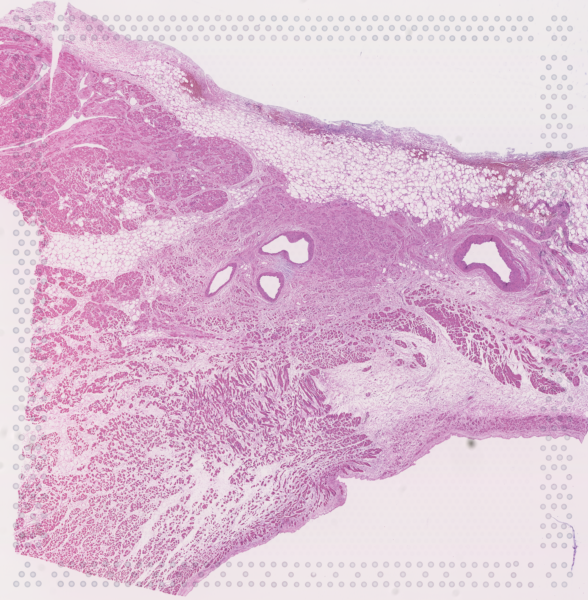
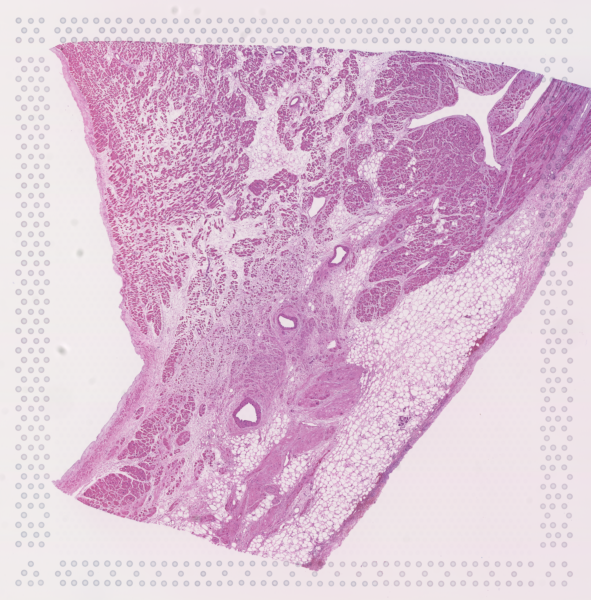
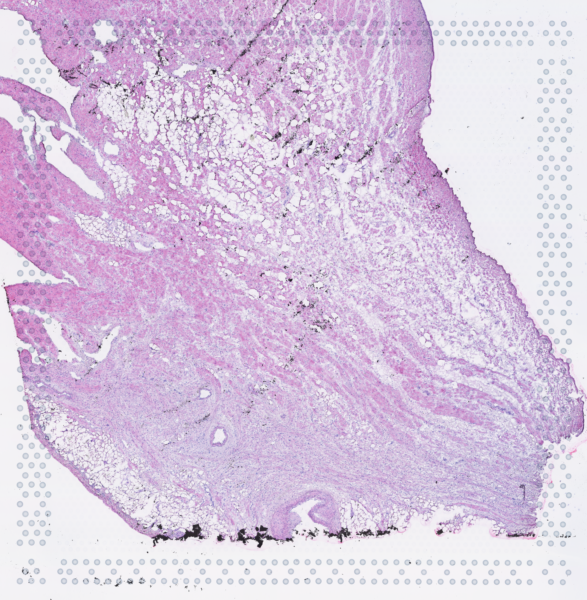
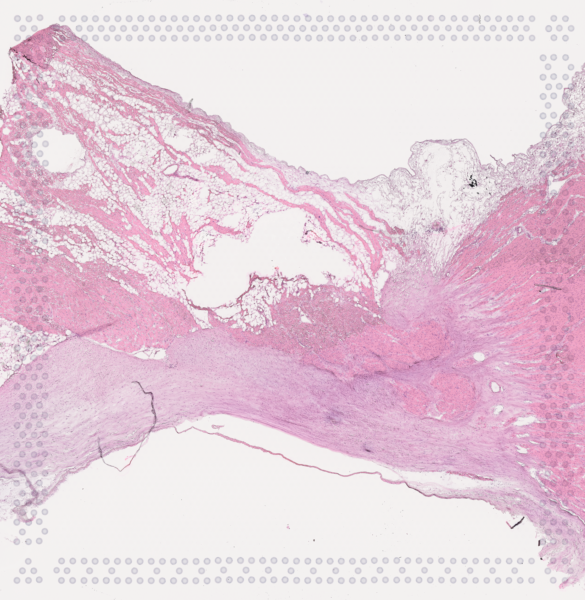
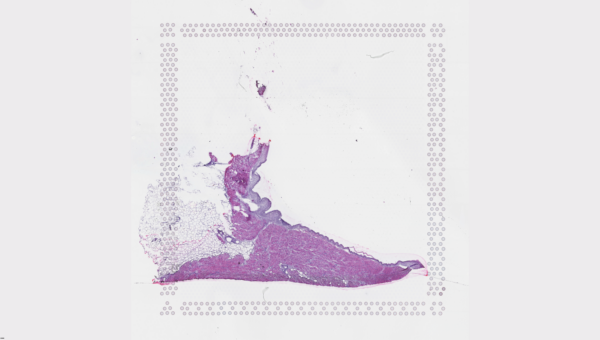
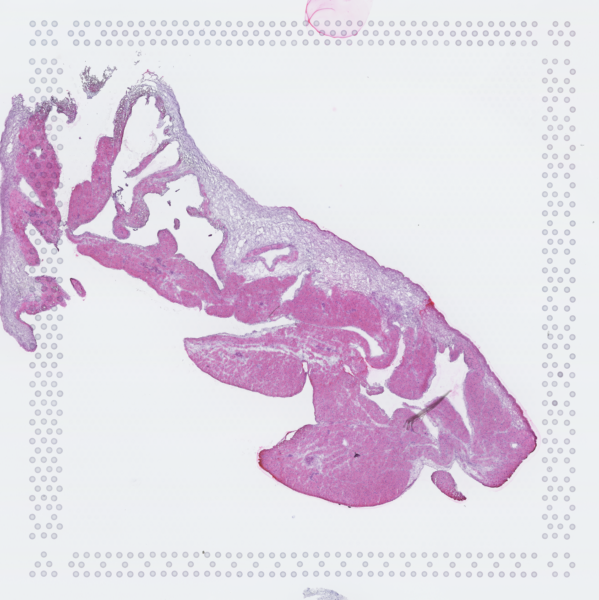
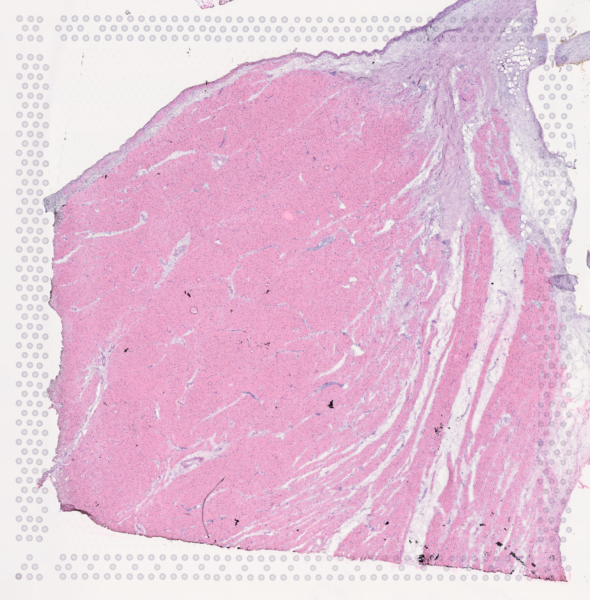
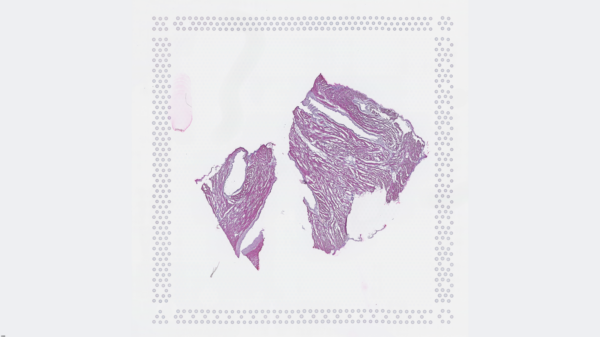
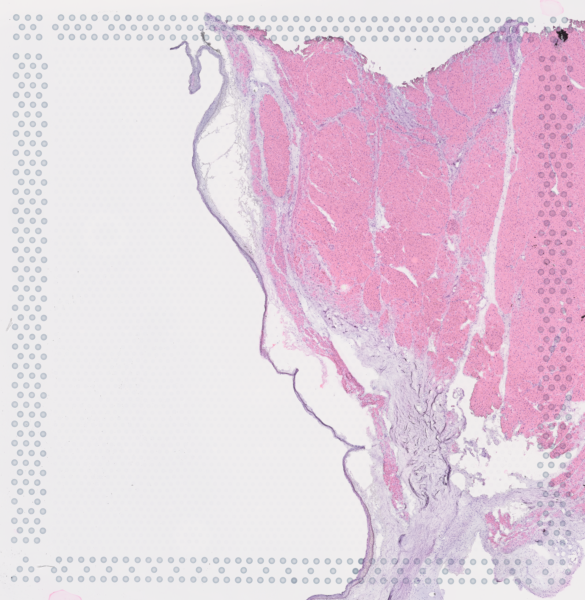
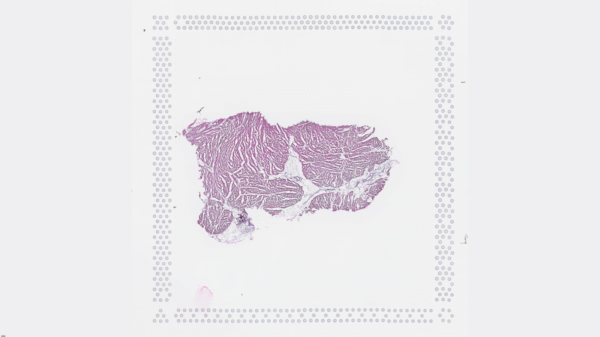
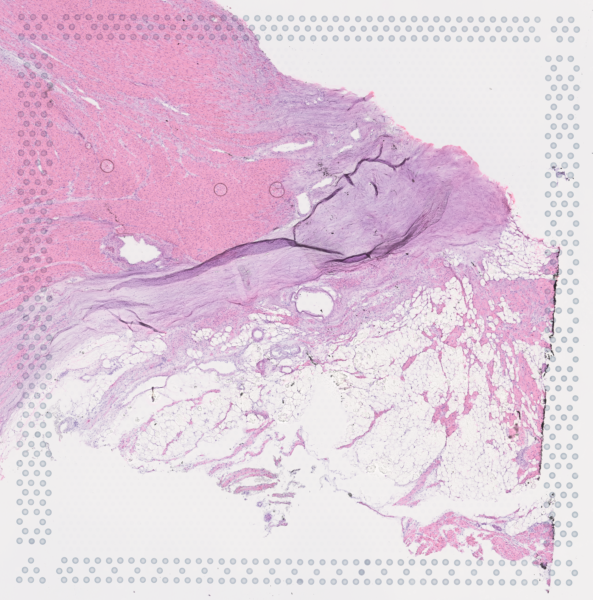
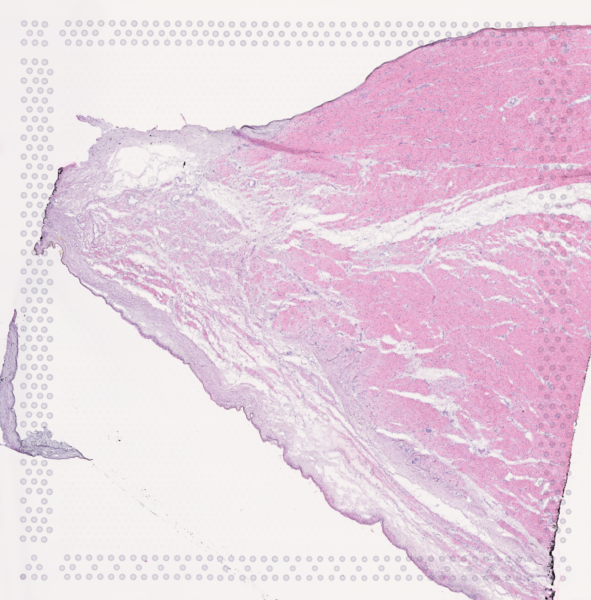
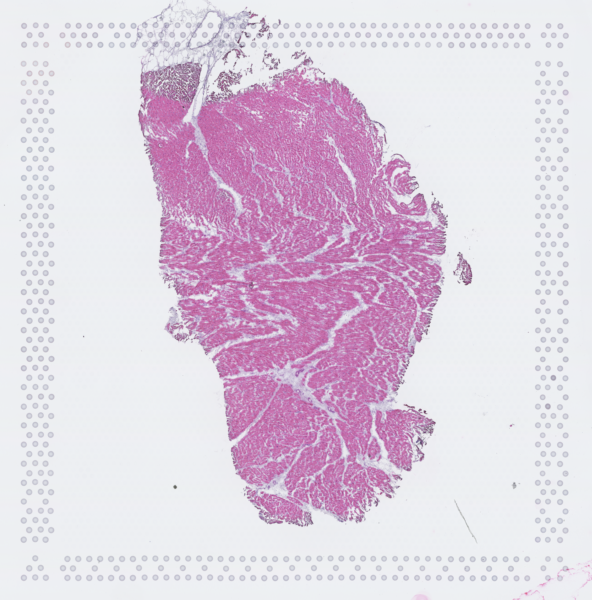
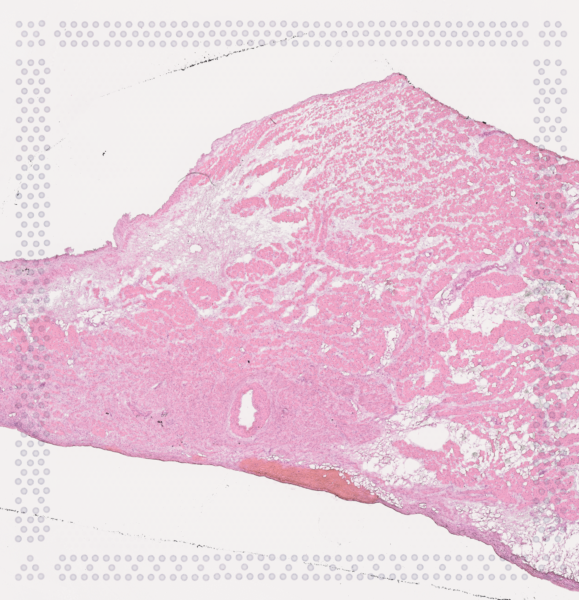
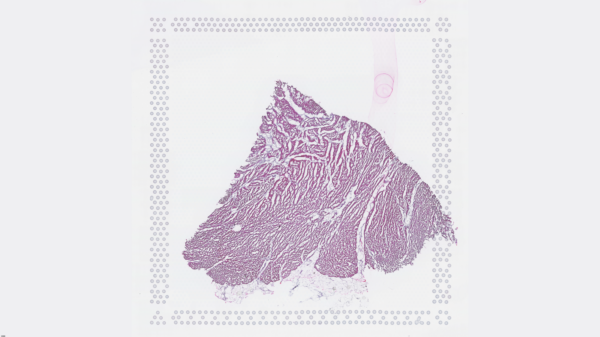
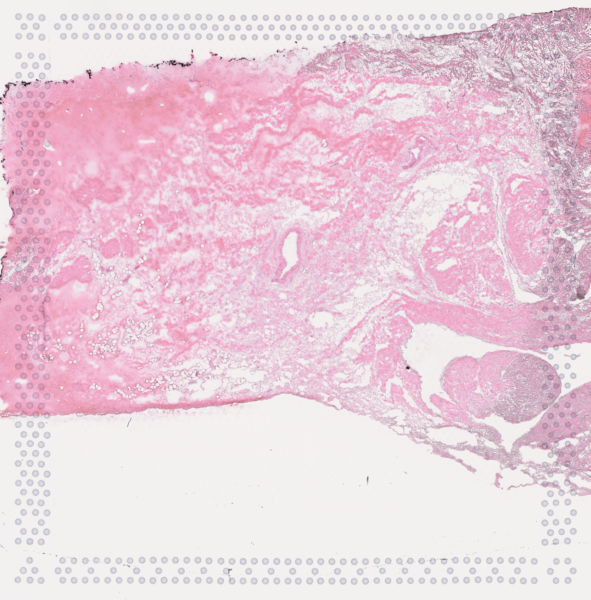
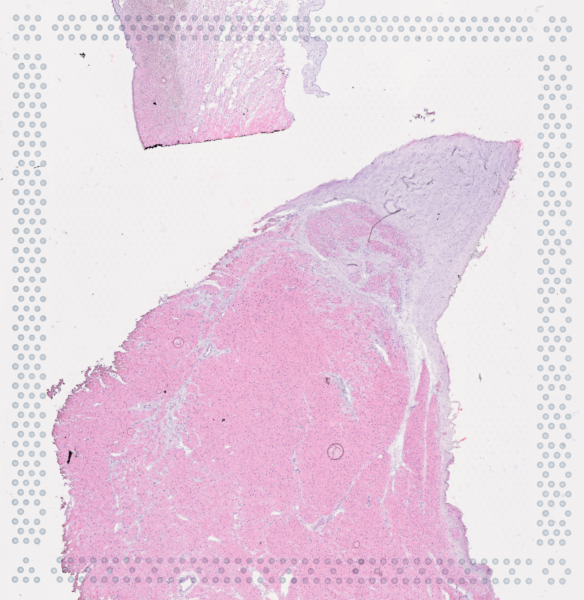
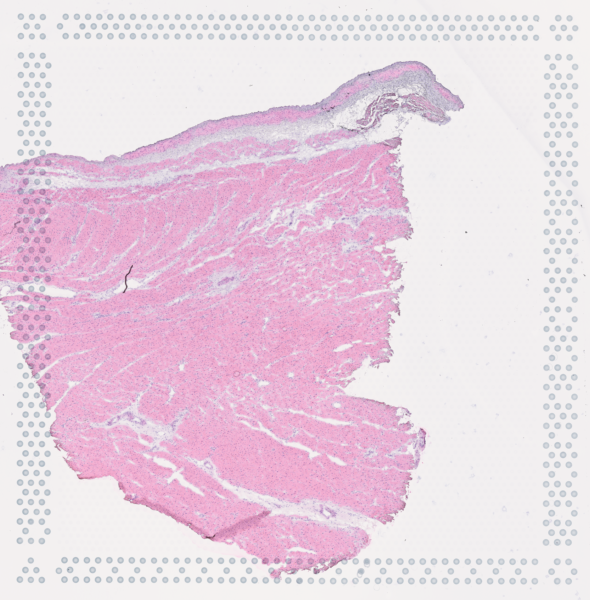
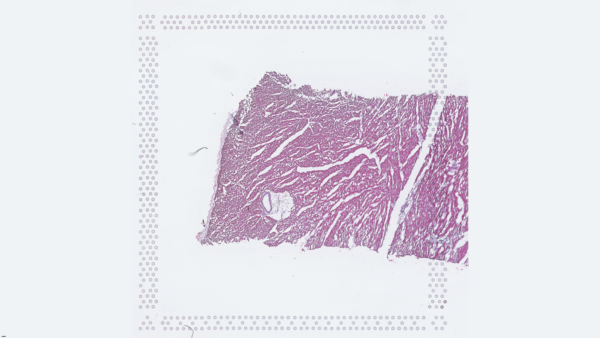
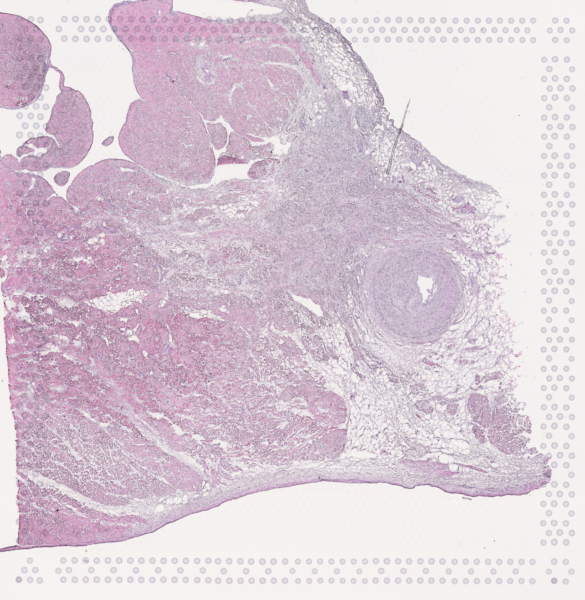
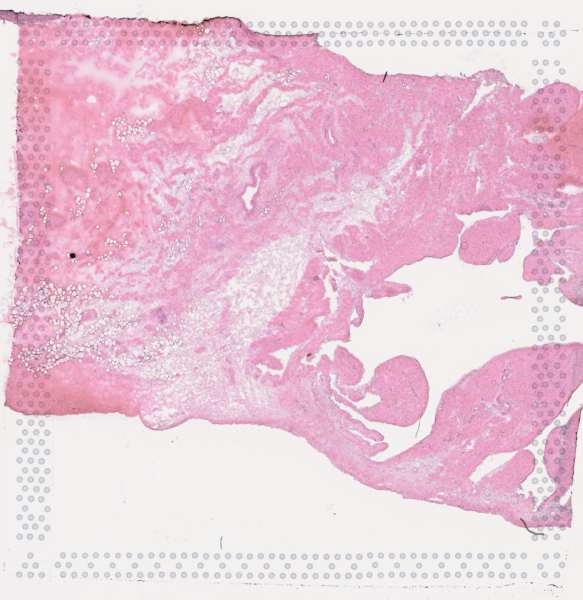
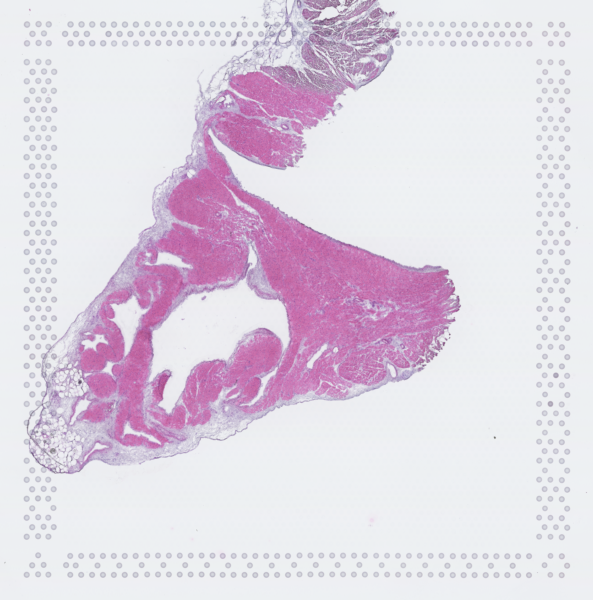
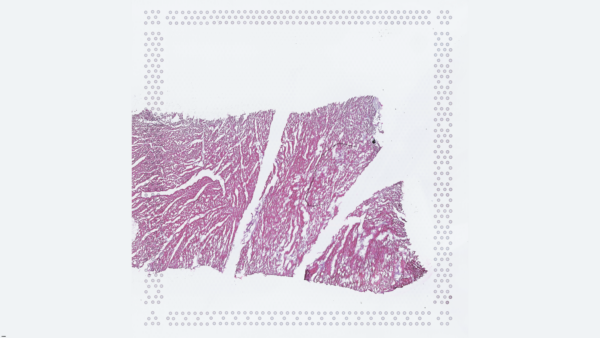
The cardiac conduction system (CCS), responsible for the regular and coordinated electrical activation of the heart, contains structures including the sinoatrial and atrioventricular nodes, atrioventricular bundle (AV bundle) and His-Purkinje network which are each home to cells with unique electrophysiological properties. We combined targeted dissection and histology to generate full-breadth transcriptomic profiles of human CCS cells for the first time.
Identified CCS cells:
- Pacemaker cell in SAN (SAN_P_cell)
- Pacemaker cell in AVN (AVN_P_cell)
- AV bundle cell
- Purkinje cell
Using cell2location, we confirmed that the identified CCS cells localised to the histologically-identified CCS structures annotated based on H&E images by an expert anatomist.
Data
View data processing and analysis code on ![]() GitHub.
GitHub.
-
chevron_rightSingle-cell/nuclei RNA and ATAC-seq
Summary Interactive view H5AD (log-normalised) H5AD (raw) CellTypist model ATAC Peak Matrix ATAC Fragments Heart Global pageview file_download file_download file_download file_download file_download Heart SAN atrial cardiomyocytes pageview file_download file_download Heart AVN atrial cardiomyocytes pageview file_download file_download -
chevron_rightVisium - H5AD
Dataset H5AD (log-normalised) H5AD (raw) left ventricular free walls (LV) file_download file_download right ventricular free walls (RV) file_download file_download left atria (LA) file_download file_download right atria (RA) file_download file_download apex (AX) file_download file_download interventricular septum (SP) file_download file_download sino-atrial node (SAN) file_download FFPE-SAN file_download OCT-SAN file_download FFPE-SAN file_download OCT-SAN atrioventricular node (AVN) file_download file_download -
chevron_rightVisium - Interactive sections with cell2location prediction minimal abundances (q05cell_abundance)